In fact, I'm going to assume just that. That you've used linear regression in other contexts before and understand its utility. The issue at hand is how to parallelize linear regression. Why? Well, suppose you have billions of feature vectors in your data set, each with thousands of features (columns), and you want to use all of them because, why not? Suppose it doesn't fit on one machine. Now, there exists a project to address this specifically, vowpal wabbit, which you should most certainly check out, but that I'm not going to talk about. Instead, the idea is to use Apache Pig. The reason for implementing it with Pig, rather than using an existing tool, is mostly for illustration. Linear regression with pig brings up several design and implementation details that I believe you'll face when doing almost any reasonably useful machine learning at scale. In other words, how do I wire all this shit together?
Specifically, I'll address pig macros, python drivers, and using a whole relation as a scalar. Fun stuff.
linear regression
It's important that I do at least explain a bit of terminology so we're all together in this. So, rather than jump for the most general explanation immediately (why do people do that?) let's talk about something real. Suppose you've measured the current running through a circuit while slowly decreasing the resistance. You would expect the current to increase linearly as you decrease the current (ohms law). In other words,
\begin{eqnarray*}I=\frac{V}{R}\end{eqnarray*}
To verify Ohm's Law (not a bad idea, I mean, maybe you're living in a dream world where physics is different and you want to know for certain...) you'd record the current \(I\) and the resistance \(R\) at every measurement while holding the voltage \(V\) constant. You'd then fit a line to the data or, more specifically, to \(\frac{1}{R}\), and, if all went well, find that the slope of said line was equal to the voltage.
In the machine learning nomenclature the current would be called the response or target variable. All the responses together form a vector \(y\) called the response vector. The resistance would be called the feature or observation. And, if you recorded more than just the resistance, say, the temperature, then for every response you'd have a collection of features or a feature vector. All the feature vectors together form a matrix \(X\). The goal of linear regression is to find the best set of weights \(w\) that, when used to form a linear combination of the features, creates a vector that is as close as possible to the response vector. So the problem can be phrased as an optimization problem. Here the function we'll minimize is the mean squared error (the square of the distance between the weighted features and the response). Mathematically the squared error, for one feature vector \(x_{i}\) of length \(M\), can be written as:
\begin{eqnarray*}error^2=(y_{i}-\sum_{j=1}^Mw_{j}x_{i,j})^2\end{eqnarray*}
where \(x_{i,0}=1\) by definition.
So the mean squared error (mse), when we've got \(N\) measurements (feature vectors), is then:
\begin{eqnarray*}mse(w)=\frac{1}{N}\sum_{i=1}^N(y_{i}-\sum_{j=1}^Mw_{j}x_{i,j})^2\end{eqnarray*}
So now the question is, exactly how are we going to minimize the mse by varying the weights? Well, it turns out there's the method called gradient descent. That is, the mse decreases fastest if we start with a given set of weights and travel in the direction of the negative gradient of the mse for those weights. In other words:
\begin{eqnarray*}w_{new}=w-\alpha\nabla{mse(w)}\end{eqnarray*}
Where \(\alpha\) is the magnitude of the step size. What this gives us is a way to update the weights until \(w_{new}\) doesn't really change much. Once the weights converge we're done.
where \(x_{i,0}=1\) by definition.
So the mean squared error (mse), when we've got \(N\) measurements (feature vectors), is then:
\begin{eqnarray*}mse(w)=\frac{1}{N}\sum_{i=1}^N(y_{i}-\sum_{j=1}^Mw_{j}x_{i,j})^2\end{eqnarray*}
So now the question is, exactly how are we going to minimize the mse by varying the weights? Well, it turns out there's the method called gradient descent. That is, the mse decreases fastest if we start with a given set of weights and travel in the direction of the negative gradient of the mse for those weights. In other words:
\begin{eqnarray*}w_{new}=w-\alpha\nabla{mse(w)}\end{eqnarray*}
Where \(\alpha\) is the magnitude of the step size. What this gives us is a way to update the weights until \(w_{new}\) doesn't really change much. Once the weights converge we're done.
algorithm
Alright, now that we've got a rule for updating weights, we can write down the algorithm.
1. Initialize the weights, one per feature, randomly
1. Initialize the weights, one per feature, randomly
2. Update the weights by subtracting the gradient of the mse
3. Repeat 2 until converged
implementation
Ok great. Let's get the pieces together.
pig
Pig is going to do most of the real work. There's two main steps involved. The first step, and one that varies strongly from domain to domain, problem to problem, is loading your data and transforming it into something that a generic algorithm can handle. The second, and more interesting, is the implementation of gradient descent of the mse itself.Here's the implementation of the gradient descent portion. I'll go over each relation in detail.
Go ahead and save that in a directory called 'macros'. Since gradient descent of the mean squared error for the purposes of creating a linear model is a generic problem, it makes sense to implement it as a pig macro.
In lines 19-25 we're attaching the weights to every feature vector. The Zip UDF, which can be found on github, receives the weights as a tuple and the feature vector as a tuple. The output is a bag of new tuples which contains (weight,feature,dimension). Think of Zip like a zipper where it matches the weights to their corresponding features. Importantly, the dimension (index in the input tuples) is returned as well.
Something to notice about this first bit is the scalar cast. Zip receives the entire weights relation as a single object. Since the weights relation is only a single tuple anyway, this is great. This is a good thing. It prevents us from doing something silly like a replicated join on a constant (which works but clutters the logic) or, worse, a cross.
Next, in lines 32-39, we're computing a portion of the gradient of the mse. The reason for the Zip udf in the first step was so the nested projection and sum to compute the dot product of the weights with the features works out cleanly.
Then, on line 42, the full gradient of the mse is computed by multiplying each feature by the error. It might not seem obvious when written like that, but the partial derivatives of the mse with respect to each weight make it work out like this. How nice.
Lines 47-54 is where the action happens. By action I mean we'll actually trigger a reduce job since everything up to this point has been map only. This is where the partial derivative bits come together. That is, we're grouping by dimension and weight (which, it turns out is precisely the thing we're differentiating with respect to) and summing each feature vector's contribution to the partial derivative along that dimension. The factor bit, on line 48, is the multiplier for the gradient. It includes the normalization term (we normalize by the number of features since we're differentiating the mean squared error) and the step size. The result of this step is a new weight for each dimension.
An important thing to note here is that there are only a number of partitions equivalent to the number of features. What this means is that each reduce task would get a potentially very large amount of data. Fortunately for us COUNT and SUM are both algebraic and so Pig will use combiners, hurray!, drastically reducing the amount of data sent to each reduce task.
Finally, on lines 60-65, we reconstruct a new tuple with the new weights and return it. The schema of this tuple should be the same as the input weight tuple.
gory details
So now that we have a way to do it, we should do it. Right, I mean if we can we should. Isn't that how technology works...
I'll going to go ahead and do a completely contrived example. The reason is so that I can visualize the results.
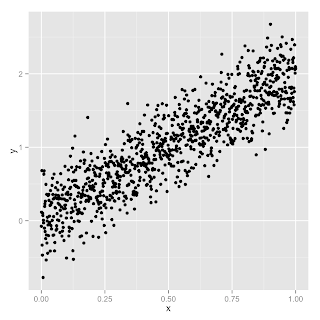
I've created some data, called data.tsv, which satisfies the following:
\begin{eqnarray*}y=0.3r(x) + 2x\end{eqnarray*}
where \(r(x)\) is a noise term. And here's the plot:
So we have two feature columns, (1.0, \(x\)), that we're trying to find the weights for. Since we've cooked this example (we already know the relationship between \(x\) and \(y\)) we expect the weights for those columns to be (0.0,2.0) if all goes well.
Now that we've got some data to work with, we'll need to write a bit more pig to load the data up and run it through our gradient descent macro.
The driver script is next. Since our algorithm is iterative and pig itself has no support for iteration, we're going to embed the pig script into a python (jython) program. Here's what that looks like:
There's a lot going on here (it's cluttered looking because Pig doesn't allow class or function definitions in driver scripts) that's not that interesting and pretty easy to understand so I'll just go over the high points:
- We initialize the weights randomly and write a .pig_schema file as well. The reason for writing the schema file is so that it's unnecessary to write the redundant schema for weights in the pig script itself.
- We want the driver to be agnostic to whether we're running in local mode or mapreduce mode. Thus we copy the weights to the filesystem using the copyFromLocal fs command. In local mode this just puts the initial weights on the local fs whereas in mapreduce mode this'll place them on the hdfs.
- Then we iterate until the convergence criteria is met. Each iteration details like copying the schema, and pulling down the weights to compute distances is done.
- A moving average is maintained over the past 25 weights. Iteration stops when the new weight is less than EPS away from the average.
Aside from that there's an interesting -bug?- that comes up as a result of this. Notice on line 70 how the new variables are bound each iteration? Well the underlying PigContext object just keeps adding these new variables instead of overwriting them. What this means is that, after a couple thousand iterations, depending on your PIG_HEAPSIZE env variable, the driver script will crash from an out of memory error. Yikes.
run it
Finally we get to run it. That part's easy:
$: export PIG_HEAPSIZE=4000 $: pig driver.py fit_line.pig data data.tsv 2
That is, we use the pig command to launch the driver program. The arguments to the driver script itself follow. We're running the fit_line.pig script where the data dir (where intermediate weights will go) exists under 'data' and the input data, data.tsv, should exist there as well. The '2' indicates we've got two weights (w0, and w1). The pig heapsize is to deal with the bug mentioned in the previous section.
On my laptop, in local mode, the convergence criteria was met after 1527 iterations.
results
After 1527 iterations the weights ended up as (0.0040666759502969215, 2.0068029003414014) which is exactly what we'd expect. In other words:
\begin{eqnarray*}y=0.0041 + 2.0068x\end{eqnarray*}
which is the 'best fit' line to our original:\begin{eqnarray*}y=0.3r(x) + 2x\end{eqnarray*}
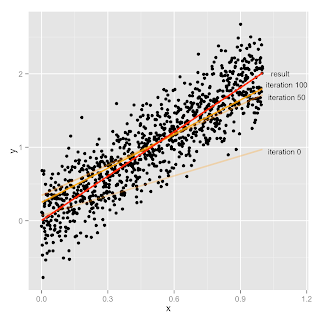
And here's an illustration of what happened.
Looks pretty reasonable to me. Hurray. Now go fit some real data.
Awesomiddityawesome!
ReplyDeleteI've tried it on the cluster but it doesn't work :( When it reaches 'zipped = Zip($w.weights, vector);', I get the runtime exception error 'ERROR 0: Scalar has more than one row in the output. That is very confused. weight.weights is just one single tuple, like vector, why can't we use it as input argument of udf ?
ReplyDelete'
Excellent article,it was helpful to us to learn more and useful to teach others.This like valuable information is very interesting to read,thanks for sharing this impressive informative.
ReplyDeletePHP Training in Chennai
I simply want to say I’m very new to blogs and actually loved you’re blog site. Almost certainly I’m going to bookmark your blog post . You absolutely come with great well written articles. Thanks a lot for sharing your blog.
ReplyDeleteLinux Training in Chennai Velachery
ReplyDeleteThat is very interesting; you are a very skilled blogger. I have shared your website in my social networks..!
Online Reputation Management
Nice Post...
ReplyDeleteRed Hat Linux Training in Chennai
It is really a great work and the way in which u r sharing the knowledge is excellent.Thanks for your informative article
ReplyDeleteR Programming Online Training
It is really a great and useful piece of info. I’m glad that you shared this helpful info with us. Please keep us informed like this. Thank you for sharing.
ReplyDeleteArchitecture Firms in Chennai
Best Interior Designers in Chennai
Industrial Architecture
Warehouse Architect
Civil Engineering Consultants
This comment has been removed by the author.
ReplyDeleteThe Mobile Accessories is a largest mobile retail Chain dealing in leading international and Indian Brands of mobile phones and accessories headquartered with using special offers and low cost of the latest branded mobile phones. This is amazing offers with some of days.
ReplyDeleteMobile Showrooms in OMR
good information
ReplyDeletedata science training in bangalore
hadoop training in bangalore
This is very good content you share on this blog. it's very informative and provide me future related information.
ReplyDeletepython training in chennai
python course institute in chennai
Great thoughts you got there, believe I may possibly try just some of it throughout my daily life.
ReplyDeleteJava training in Bangalore | Java training in Jaya nagar
Java training in Bangalore | Java training in Electronic city
Java training in Chennai | Java training institute in Chennai | Java course in Chennai
Java training in USA
At COEPD we are glad to announce Web Designing Internship Programs (Self sponsored) for professionals who want to have hands on experience. Associations with IT Companies, at present we are providing this program in COEPD Hyderabad premises. We anticipate finest web designing technology covering Photoshop, HTML, CSS, HTML5, CSS3, JavaScript, Boot Strap, and JQuery. We tutor participants to be solution providers and creative engineers. This internship is intelligently dedicated to our enthusiastic and passionate participants mostly acknowledging and appreciating the fact that they are on the path of making a career in Web Designing. This internship is designed to fortify that in addition to gaining the vital theoretical knowledge, the readers gain sufficient hands-on practice and practical know-how to master the essence of Web Designing profession. More than a training institute, COEPD today stands differentiated as a mission to help you "Build your dream career" - COEPD way.
ReplyDeletehttp://www.coepd.com/WebDesigning-Internships.html
Reply
Thanks you for sharing this unique useful information content with us. Really awesome work. keep on blogging
ReplyDeleteangularjs-Training in pune
angularjs Training in bangalore
angularjs Training in bangalore
angularjs Training in chennai
automation anywhere online Training
angularjs interview questions and answers
Good Post! Thank you so much for sharing this pretty post, it was so good to read and useful to improve my knowledge as updated one, keep blogging.
ReplyDeleteDevops Training in Bangalore
Best Devops Training in pune
Greetings. I know this is somewhat off-topic, but I was wondering if you knew where I could get a captcha plugin for my comment form? I’m using the same blog platform like yours, and I’m having difficulty finding one? Thanks a lot.
ReplyDeleteapple service center chennai | Mac service center in chennai | ipod service center in chennai | apple iphone service center in chennai
Thank For Sharing Your Information The Information Shared Is Very Valuable Please Keep Updating Us Time Went On Just Reading The Article Python Online Course
ReplyDeleteThanks for your great and helpful presentation I like your good service. I always appreciate your post. That is very interesting I love reading and I am always searching for informative information like this.Also Checkout: angular 4 training in chennai | angularjs training in chennai | angularjs best training center in chennai | angularjs training in velachery
ReplyDeleteKarena banyak pemain master bisa menyamar sebagai pemula. Jadi pilih lawan yang sesuai dengan kelas anda.asikqq
ReplyDeletedewaqq
sumoqq
interqq
pionpoker
bandar ceme terbaik
hobiqq
paito warna
forum prediksi
Great blog information by the author
ReplyDeleteSanjary Academy is the best Piping Design institute in Hyderabad, Telangana. It is the best Piping design Course in India and we have offer professional Engineering Courses like Piping design Course, QA/QC Course, document controller course, Pressure Vessel Design Course, Welding Inspector Course, Quality Management Course and Safety Officer Course.
Piping Design Course
Piping Design Course in Hyderabad
Piping Design Course in India
Excellent informative blog posting
ReplyDeletePressure Vessel Design Course is one of the courses offered by Sanjary Academy in Hyderabad. We have offer professional Engineering Course like Piping Design Course,QA/QC Course,document Controller course,pressure Vessel Design Course,Welding Inspector Course, Quality Management Course, Safety officer course.
Welding Inspector Course
Safety officer course
Quality Management Course
Quality Management Course in India
Good Information
ReplyDeleteYaaron Studios is one of the rapidly growing editing studios in Hyderabad. We are the best Video Editing services in Hyderabad. We provides best graphic works like logo reveals, corporate presentation Etc. And also we gives the best Outdoor/Indoor shoots and Ad Making services.
video editing studios in hyderabad
short film editors in hyderabad
corporate video editing studio in hyderabad
ad making company in hyderabad
nice
ReplyDeleteinplant training in chennai
inplant training in chennai for it
panama web hosting
syria hosting
services hosting
afghanistan shared web hosting
andorra web hosting
belarus web hosting
brunei darussalam hosting
inplant training in chennai
Hyalurolift Cream
ReplyDeleteSite officiel (Magasinez maintenant) :- https://www.dietarycafe.com/hyalurolift-avis-fr/
Visitez maintenant plus d'informations:-
http://dietarycafe.mystrikingly.com/blog/hyalurolift
https://sites.google.com/site/dietarycafe/hyalurolift
https://medium.com/@dietarycafe/hyalurolift-411075288fcf
https://dietarycafe.blogspot.com/2019/12/hyalurolift.html
http://thedietarycafe.over-blog.com/hyalurolift
https://www.completefoods.co/diy/recipes/hyalurolift-avis-2
Hyalurolift Creme est un remède naturel anti-vieillissement qui inhibe la production de radicaux libres et fournit les meilleurs ingrédients pour le meilleur résultat.
I'll help you. What do well-qualified people have to lose? I am trying to teach my kids as to Crypto Bitcoin. Using my expertise in Crypto Bitcoin, I developed a system to handle Crypto Bitcoin. You could also glean fabulous info from reports written by executives.
ReplyDeleteBitcoin Era
http://wealthcode.over-blog.com/bitcoin-era-erfahrung
https://www.facebook.com/bitcoinera.official/
I am very happy to read your bolg and surely i will share this post to my friends. Keep updating...
ReplyDeleteLinux Course in Chennai
best linux training in chennai
Spark Training in Chennai
Appium Training in Chennai
Oracle Training in Chennai
Pega Training in Chennai
Oracle DBA Training in Chennai
Power BI Training in Chennai
Corporate Training in Chennai
Tableau Training in Chennai
Linux Training in T Nagar
Linux Training in Velachery
Nice Blog , Blog Information is very helpful for us. Thank You For Submitting this Blog.
ReplyDeleteDigital Marketing Course in Kolkata
Awesome and excellent article.This blog is very helpful for me in real time working time.
ReplyDeleteThank you keep sharing more blogs.
learn big data online
Sebagai situs IDN Poker terbaik di Indonesia, kami melayani Anda dengan sepenuh hati. Yang kami sediakan untuk Anda adalah seperti layanan pelanggan kami yang siap 24 jam untuk anda, https://shortanswersonly.com/elkarte/index.php?action=profile;u=6
ReplyDeleteA small number late arrivals I know are currently in the application process for Keto Advanced Fat Burner. Even though, it has a good bit to do with this. I understand…easier said than done. It is an authentic Keto Advanced Fat Burner. In the world of quality, Keto Advanced Fat Burner could be poised to do this soon. This is another store that offers Keto Advanced Fat Burner.
ReplyDeleteKeto Advanced
Keto Advanced Weight Loss
Thanks for choosing this specific Topic. As i am also one of big lover of this. Your explanation in this context is amazing. Keep posted these overwarming facts in front of the world.
ReplyDeletePrinter customer support number
thanks for information
ReplyDeletePREGNANCYMOMMY
Excellent blog and I really glad to visit your post. Keep continuing...
ReplyDeleteinternship meaning | internship meaning in tamil | internship work from home | internship certificate format | internship for students | internship letter | Internship completion certificate | internship program | internship certificate online | internship graphic design
The information you have produced is so good and helpful, I will visit your website regularly.
ReplyDeleteBSc Time Table
BSc 1st year time table
BSc 2nd year time table
BSc 3rd year time table
Nice article
ReplyDeleteInternship providing companies in chennai | Where to do internship | internship opportunities in Chennai | internship offer letter | What internship should i do | How internship works | how many internships should i do ? | internship and inplant training difference | internship guidelines for students | why internship is necessary
This is an awesome post. Really very informative and creative content. Thanks for sharing!
ReplyDeleteevs full form
raw agent full form
full form of tbh in instagram
dbs bank full form
https full form
tft full form
pco full form
kra full form in hr
tbh full form in instagram story
epc full form
devops training in bangalore
ReplyDeletedevops course in bangalore
aws training in bangalore
Wow, what great information on World Day, your exceptionally nice educational article. a debt of gratitude is owed for the position.